
They have identified a promising direction for use of metabolic imaging for macrophage classification. Editor's evaluation The authors introduce a machine learning based classifier for M1 and M2 polarised macrophages based on autofluorescence lifetime parameters excited by two-photon excitation in the NAD(P) H emission band following during uncoupling of oxidative phosphorylation. #Cellprofiler custom script full#To conclude, 2P-FLIM with the integration of machine learning models is showed to be a powerful technique for analysis of both human macrophage metabolism and polarisation at full FoV and single-cell level. Applying a random forests model, we identify three strongly governing FLIM parameters, achieving an area under the receiver operating characteristics curve (ROC-AUC) value of 0.944 and out-of-bag (OBB) error rate of 16.67% when classifying human macrophages in a full field-of-view image. The stratification and parameters emanating from a full field-of-view and single-cell FLIM approach serve as the basis for machine learning models. We uncovered FLIM parameters that are pronounced under the action of carbonyl cyanide-p-trifluoromethoxyphenylhydrazone (FCCP), which strongly stratifies the phenotype of polarised human macrophages however, this performance is impacted by donor variability when analysing the data at a single-cell level. 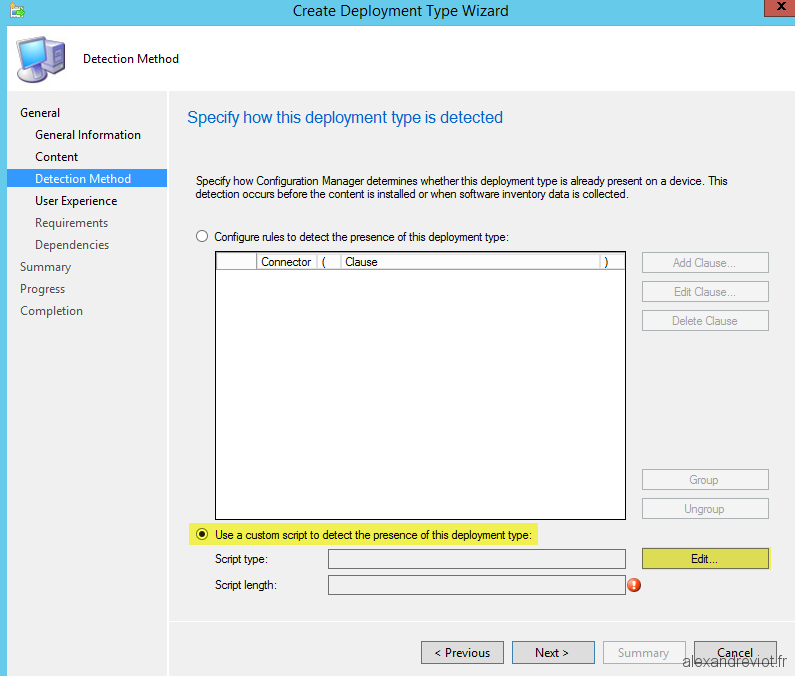
These were challenged in real time with small-molecule perturbations of metabolism during imaging. Large field-of-view images of individual polarised macrophages were obtained using fluorescence lifetime imaging microscopy (FLIM). 
Macrophages derived from human blood-circulating monocytes were polarised using established protocols and metabolically challenged using small molecules to validate their responding metabolic actions in extracel-lular acidification and oxygen consumption. In this study, we utilise fluorescence lifetime imaging of NAD(P)H-based cellular autofluorescence as a non-invasive modality to classify two contrasting states of human macrophages by proxy of their governing metabolic state. Our results distinguish Omnipose as a powerful tool for characterizing diverse and arbitrarily shaped cell types from imaging data. Finally, we demonstrate the utility of Omnipose in the characterization of extreme morphological phenotypes that arise during interbacterial antagonism. Furthermore, the benefits of Omnipose extend to non-bacterial subjects, varied imaging modalities and three-dimensional objects. We show that Omnipose achieves unprecedented segmentation performance on mixed bacterial cultures, antibiotic-treated cells and cells of elongated or branched morphology. Unique network outputs such as the gradient of the distance field allow Omnipose to accurately segment cells on which current algorithms, including its predecessor, Cellpose, produce errors. Here, we present Omnipose, a deep neural network image-segmentation algorithm. We hope these changes will make CellProfiler an even better tool for current users and will provide new users better ways to get started doing quantitative image analysis.Īdvances in microscopy hold great promise for allowing quantitative and precise measurement of morphological and molecular phenomena at the single-cell level in bacteria however, the potential of this approach is ultimately limited by the availability of methods to faithfully segment cells independent of their morphological or optical characteristics. We’ve also added more explanations to CellProfiler’s settings to help new users get started. #Cellprofiler custom script code#We’ve also made changes to CellProfiler’s underlying code to make it faster to run and easier to install, and we’ve added the ability to process images in the cloud and using neural networks (deep learning). In this release, we’ve added the capability to find and measure objects in three-dimensional (3D) images. Pipelines are easy to save, reuse, and share, helping improve scientific reproducibility. #Cellprofiler custom script download#Researchers can download an online example workflow (that is, a “pipeline”) or create their own from scratch. #Cellprofiler custom script software#The third major release of our free open-source software CellProfiler is designed to help biologists working with images, whether a few or thousands. 
Thus, many biologists find they need software to analyze images easily and accurately. Looking at the resulting images by eye would be extremely tedious, not to mention subjective. The “big-data revolution” has struck biology: it is now common for robots to prepare cell samples and take thousands of microscopy images. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
